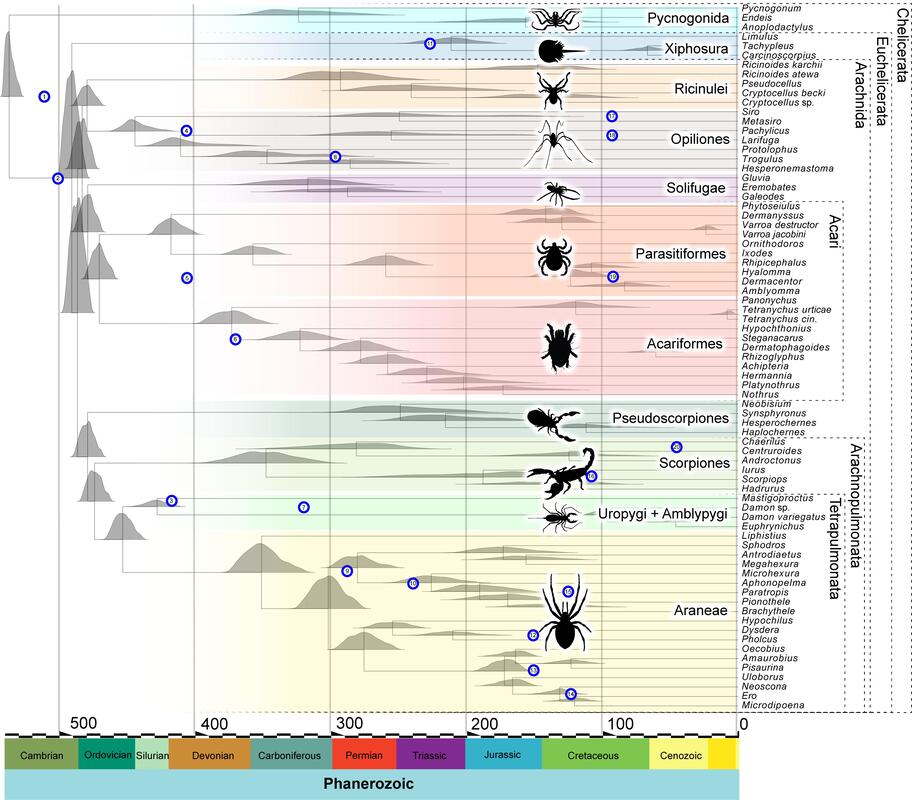
Arthropod terrestrialisation using phylogenomics, comparative genomics and total evidence dating
Animals have marine origins and only four lineages have adapted to life on land. These groups are the vertebrates (e.g. reptiles, mammals), molluscs (e.g. land snails, slugs), the onychophora (velvet worms) and the arthropods (e.g. insects, spiders). The water-to-land transition is referred as terrestrialisation, and is an extreme case of adaptation to a new environment.
Our research is focused on the arthropod lineage, and we are interested in knowing 'how' and 'when' these organisms have colonised the land. We are particularly interested in the comparative genomics of one of the groups that conform it, the chelicerates (e.g. spiders, scorpions, mites). Chelicerates contains marine and land representatives as well as abundant terrestrial fossil record. We combine the genomic and fossil information of these organisms to study the modifications that allowed marine chelicerates to adapt to life on land.
Animals have marine origins and only four lineages have adapted to life on land. These groups are the vertebrates (e.g. reptiles, mammals), molluscs (e.g. land snails, slugs), the onychophora (velvet worms) and the arthropods (e.g. insects, spiders). The water-to-land transition is referred as terrestrialisation, and is an extreme case of adaptation to a new environment.
Our research is focused on the arthropod lineage, and we are interested in knowing 'how' and 'when' these organisms have colonised the land. We are particularly interested in the comparative genomics of one of the groups that conform it, the chelicerates (e.g. spiders, scorpions, mites). Chelicerates contains marine and land representatives as well as abundant terrestrial fossil record. We combine the genomic and fossil information of these organisms to study the modifications that allowed marine chelicerates to adapt to life on land.
Tracing the origins of multicellularity in animals and fungi
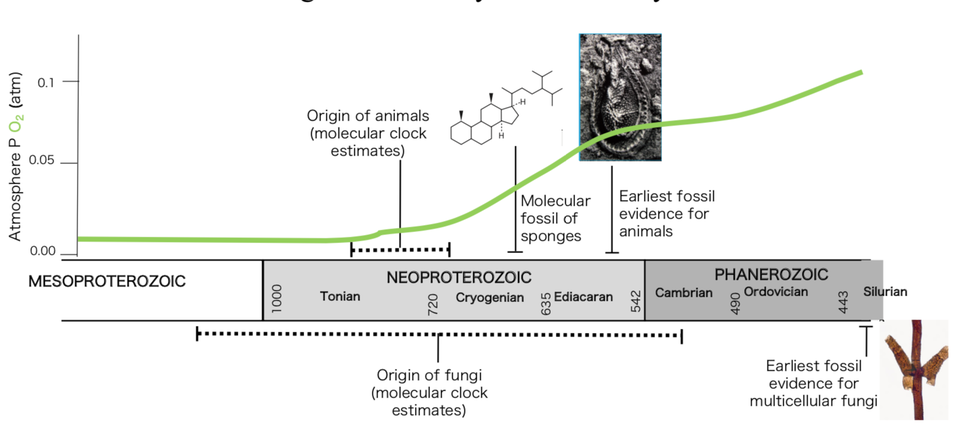
Life on Earth started around 4 billion years ago, shortly after our planet was formed, as suggested by the traces of single-cell organisms found on the fossil record. However, the origin of multicellularity was only attained during the last billion years, and represented one of the most important evolutionary leaps in the history of life. The transition to complex multicellularity has been acquired independently only in a few groups, all eukaryotes, such as plants, animals, fungi, green, brown and red algae. Two of the most iconic multicellular lineages, animals and fungi, are allied in a supergroup called Opisthokonta that also includes several unicellular relatives. Therefore, understanding this transition in both lineages, closely related organisms that share a single-cell common ancestor, offers a unique opportunity to test several hypotheses with regard to the evolution of complex life in our planet.
Life on Earth started around 4 billion years ago, shortly after our planet was formed, as suggested by the traces of single-cell organisms found on the fossil record. However, the origin of multicellularity was only attained during the last billion years, and represented one of the most important evolutionary leaps in the history of life. The transition to complex multicellularity has been acquired independently only in a few groups, all eukaryotes, such as plants, animals, fungi, green, brown and red algae. Two of the most iconic multicellular lineages, animals and fungi, are allied in a supergroup called Opisthokonta that also includes several unicellular relatives. Therefore, understanding this transition in both lineages, closely related organisms that share a single-cell common ancestor, offers a unique opportunity to test several hypotheses with regard to the evolution of complex life in our planet.
Methods in phylogenetics and molecular divergence time estimation
The advent of molecular biology and the development of the molecular clock theory allowed the use of genomes as a second record of life. This new source of evidence has been added to classical way of study the evolutionary relationships, through the study of morphology, in which early phylogenetic studies mostly relied on. Moreover, by including morphological data, fossil morphology could be used as a phylogenetical character. The integration of fossil data into a phylogenies with living species has become both a theoretical and computational challenge. The phylogenetic approaches for including phenotypic data in such analysis has had modest progress during last decades. New empirical studies are needed to test the different methods, both in simulated and empirical data.
In our research group we study the methods of implementing different phylogenetic signals, in order to use the most comprehensive information. Furthermore, we are interested in assessing the performance of different models in molecular dating analysis.
The advent of molecular biology and the development of the molecular clock theory allowed the use of genomes as a second record of life. This new source of evidence has been added to classical way of study the evolutionary relationships, through the study of morphology, in which early phylogenetic studies mostly relied on. Moreover, by including morphological data, fossil morphology could be used as a phylogenetical character. The integration of fossil data into a phylogenies with living species has become both a theoretical and computational challenge. The phylogenetic approaches for including phenotypic data in such analysis has had modest progress during last decades. New empirical studies are needed to test the different methods, both in simulated and empirical data.
In our research group we study the methods of implementing different phylogenetic signals, in order to use the most comprehensive information. Furthermore, we are interested in assessing the performance of different models in molecular dating analysis.